

Paradigm Shift Intervention Monitoring
 twitter
twitter

Commentary
Lack of
Sero-typing of US Flu A Positives Raises trH3N2 Concerns
Recombinomics Commentary 16:11
November 11, 2011
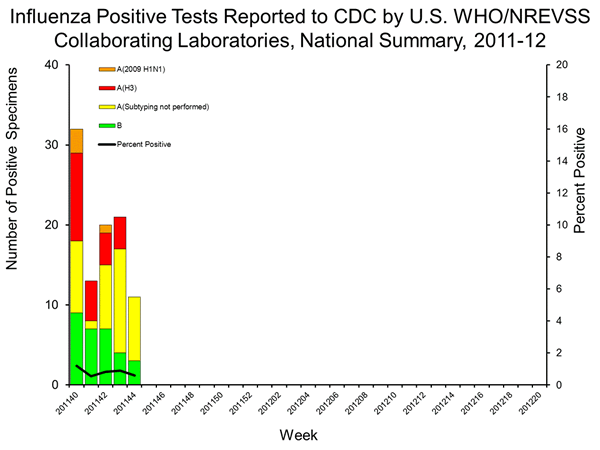
Detection of trH3N2 requires sub-typing because state labs do not have PCR tests for trH3N2. Detection requires an influenza A positive, followed by a serotype testing showing low or no activity for seasonal H3. These samples (influenza A positive, H3 negative) are then sent to the CDC for trH3N2 confirmation, so samples that are not sub-typed are not sent.
Moreover, the samples tested are also influenced by the CDC early release MMWR report on trH3N2 in July and August. The two case described has a novel constellation of flu genes, which included an M gene from H1N1pdm09. The first case (2M) represented by A/Indiana/08/2011 had no swine contact supporting human to human transmission. Similar transmission was support by the two other trH3N2 cases confirmed in 2011.
One, A/Pennsylvania/40/2010, was initially typed as seasonal H3N2. Reporting was delayed 5 months and the sequence was released more than 6 months after infection (on a Sunday at GISAID). The sequence of all 8 gene segments match another 2010 isolate, A/Wisconsin/12/2010, signaling human transmission.
Human transmission was lab confirmed in the second case reported in 2011, the daughter of the Minnesota index case, represented by A/Minnesota/11/2010. The daughter was trH3N2 lab confirmed (serologically,) and other symptomatic family members tested “inconclusive”, but were also likely infected by the same trH3N2.
The H3 from all three of these 2010 isolates were closely related to each other, as well as the first two reported cases due to 2011 infections. The detection of these two cases (A/Indiana/08/2011 and A/Pennsylvania/09/2011) was followed by two more cases in Pennsylvania, A/Pennsylvania/10/2011 and A/Pennsylvania/11/2011, which were virtually identical to the Indiana H3 sequence.
Thus, at the time of the MMWR early release there was ample evidence for human transmission of the novel trH3N2, yet the CDC limited requests to samples from cases with a swine exposure. This heavy bias on samples tested for trH3N2 is extended in the above figure, which reflects a lack of sub-typing, reflecting a prioritizing of testing of samples with a swine contact.
Three more trH3N2 cases were identified from October collections and all had a loose swine exposure history, although none of the swine have been reported as symptomatic, including those evaluated by the Indiana State Board Of Animal Health (BOAH) because the second Indiana case, A/Indiana/10/2011, was a veterinarian (59M).
Thus, although none of the 7 confirmed trH3N2 have been linked to exposure to symptomatic swine, or swine with an influenza infection. has been reported, and no swine isolate with the constellation of flu genes seen in the human trH3N2 cases has been found, the CDC continues to target testing of samples with a swine exposure, while ignoring or delaying tests on samples with no such "exposure".
This limited testing continues to be hazardous to the world’s health.
A request for samples from adolescents with flu-like symptoms, like those atypical pneumonia cases in Shelby County, Indiana, is long overdue.
Recombinomics
Presentations
Recombinomics
Publications
Recombinomics
Paper
at Nature Precedings